Haplotype Blocks and Tagging SNPs
advertisement

Association Studies, Haplotype
Blocks and Tagging SNPs
Prof. Sorin Istrail
Association studies
Disease
Responder
Allele 0
Control
Non-responder
Allele 1
Marker A:
Allele 0 =
Allele 1 =
Marker A is
associated with
Phenotype
Association studies
• Evaluate whether nucleotide
polymorphisms associate with
phenotype
A
C
G
A
G
A
C
G
A
T
A
T
A
A
G
C
T
A
G
T
A
T
G
G
T
A
T
G
G
G
Association studies
A
C
G
A
G
A
C
G
A
T
A
T
A
A
G
C
T
A
G
T
A
T
G
G
T
A
T
G
G
G
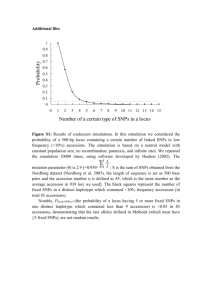
Hypothesis – Haplotype Blocks?
The
genome consists largely of blocks of
common SNPs with relatively little recombination
within the blocks
Patil et al., Science, 2001;
Jeffreys et al., Nature Genetics, 2001;
Daly et al., Nature Genetics, 2001
Haplotype Block Structure
LD-Blocks, and 4-Gamete Test Blocks
200 kb
Sense genes
DNA
Antisense genes
SNPs
Haplotype
blocks
1
2
3
4
One definition of block
• Based on the Four Gamete test.
• Intuition: when between two SNPs there
are all four gametes, there is a
recombination point somewhere
inbetween the two sites
Four Gamete Block Test
• Hudson and Kaplan 1985
A segment of SNPs is a block if between every pair of SNPs at
most 3 out of the 4 gametes (00, 01,10,11) are observed.
0
0
1
1
0
1
1
1
BLOCK
1
1
0
1
0
0
1
1
0
1
1
0
1
1
0
1
VIOLATES THE BLOCK DEFINITION
Finding Recombination Hotspots:
Many Possible Partitions into Blocks
A
G
A
G
A
A
C
T
C
T
C
C
T
T
T
T
T
T
A
C
C
A
C
A
G
G
T
T
T
G
A
A
A
A
A
C
T
C
T
C
T
T
All four gametes are present:
A
A
G
G
A
G
G
A
A
A
G
G
C
C
T
C
T
C
C
A
C
A
A
A
T
T
G
T
T
T
The final result is a minimum-size set
of sites crossing all constraints.
A C T A G A T A G C C T
GFind
T the
T left-most
C G A right
C Aendpoint
A C of
A T
AEliminate
C
T
C
T
A
T
G
A
T
C
G
any
constraints
crossing
any constraint
and mark the
site
Repeat
until
all
constraints
are
gone.
G Tbefore
T Ait aTrecombination
A
C
G
A
C
A
T
that site.
site.
A C T C T A T A G T A T
A C T A G C T G G C A T
Tagging SNPs
ACGATCGATCATGAT
GGTGATTGCATCGAT
ACGATCGGGCTTCCG
ACGATCGGCATCCCG
GGTGATTATCATGAT
An example of real data set
and its haplotype block
structure. Colors refer to the
founding population, one
color for each founding
haplotype
Only 4 SNPs are needed to tag
all the different haplotypes
A------A---TG-G------G---CG-A------G---TC-A------G---CC-G------A---TG--
Informativeness
A measure for the “information” a SNP contains
about about another SNP. Useful for designing SNPs Arrays
and Tagging SNPs selection.
s
h
1
0
0
1
1
0
h
2
0
0
1
0
1
Informativeness
0
0
1
1
0
0
0
1
0
1
0
1
0
0
0
1
1
0
1
1
s1
s2
s3
s4
s5
I(s1,s2) = 2/4 = 1/2
Informativeness
0
0
1
1
0
0
0
1
0
1
0
1
0
0
0
1
1
0
1
1
s1
s2
s3
s4
s5
I({s1,s2}, s4) = 3/4
Informativeness
0
0
1
1
0
0
0
1
0
1
I({s3,s4},{s1,s2,s5})
=3
0
1
0
0
0
S={s3,s4} is a
1
s1
1
s2
0
s3
1
s4
1
s5
Minimal Informative Subset
e
6
Informativeness
Graph theory insight
s
e
5
Minimum Set Cover
s
4
=
Minimum Informative Subset
s
s
1
s
2
s
3
s
4
s
5
0
0
1
1
0
0
0
0
1
1
0
0
0
0
1
1
0
1
1
1
s
s
e
5
4
3
e
3
2
e
1
e
SNPs
Edges
2
1
e
6
Informativeness
Graph theory insight
s
e
5
Minimum Set Cover {s3, s4}
s
4
=
Minimum Informative Subset
s
s
1
s
2
s
3
s
4
s
5
0
0
1
1
0
0
0
0
1
1
1
1
0
0
0
0
1
1
0
1
s
s
e
5
4
3
e
3
2
e
1
e
SNPs
Edges
2
1